 motor co-dependence mechanisms
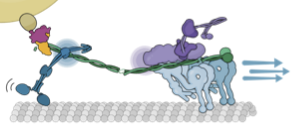
motor co-dependence mechanisms
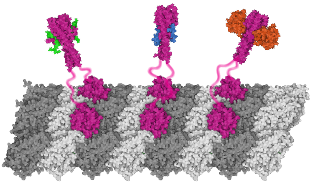
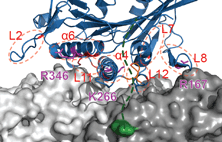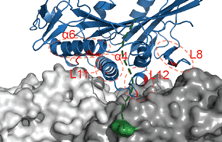
We show that complexes of dynein, KIF1C, dynactin and cargo adapter Hook3 undertake plus and minus
end-directed motility along microtubules. Both motors activate each other through the steric disinhibition
of the cargo adapter. In addition, KIF1C extends dynein-mediated runs by acting as a weak microtubule tether.
This study reveals a mechanistic understanding of co-dependence of opposite polarity motors.
Abid Ali, F., Zwetsloot, A.J., Stone, C.E., Morgan, T.E., Wademan, R.F., Carter, A.P.* and Straube, A.* (2025)
KIF1C activates and extends dynein movement through the FHF cargo adaptor
Nature Structural and Molecular Biology, doi: 10.1038/s41594-024-01418-z
[Link to open access paper]
PREPRINT bioRxiv 2023.10.26.564242v1
[Link to preprint]
 surfing the end
surfing the end
We describe an analytical mathematical model that demonstrates how processive motility is enabled by preferential
binding and unbinding of cargo-substrate links. Thereby a cargo can track newly generated or progressively disappearing
binding sites such as the tip of growing or shrinking microtubules.
Mosby, L.S., Straube, A. & Polin, M. (2023)
A general model for the motion of multivalent cargo interacting with substrates.
J. R. Soc. Interface 20: 20230510.
[Link to paper]
 weak motors cause neuronal disease
weak motors cause neuronal disease
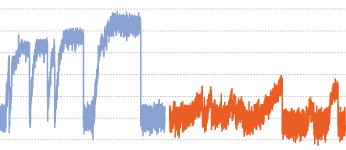
We report that KIF1C is a strong processive motor with a stall force of 5.7 pN and characterise two pathogenic
mutations P176L and R169W that cause hereditary spastic paraplegia and cerebellar dysfunction. Both mutations
reduce microtubule binding, but are fast and processive motors in single molecule assays. However, their force
generation is severely impaired and they cannot efficiently relocate cellular cargo even if working in teams.
Siddiqui, N., Roth, D., Toleikis, A., Zwetsloot, A.J., Cross, R.A. & Straube, A. (2022)
Force generation of KIF1C is impaired by pathogenic mutations
Current Biology 32(17): 3862-3870.e6
[Link to paper]
DESPATCH by Will Hancock in the same issue [Link to despatch]
SSRN 10.2139/ssrn.4075230 [Link to sneak peak]
PREPRINT bioRxiv 10.1101/2021.06.30.450611 [Link to bioRxiv preprint]
 end binding proteins
end binding proteins
Our contribution to the third edition of the Encyclopedia of Biological Chemistry is a further reading section on microtubule plus and
minus end binding proteins.
Mosby, L.S. & Straube, A. (2021)
Further Reading | Microtubule Plus and Minus End Binding Proteins
Encyclopedia of Biological Chemistry III (Third Edition), Elsevier, pages 554-566,ISBN 9780128220405,
[Link]
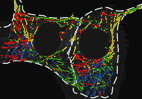
 endometrial pericyte migration
endometrial pericyte migration
A collaboration with clinical scientists interested in endometrial mesenchymal stem cells. Here we show that vascular edhesion protein-1 (VAP-1) is
required for proper cell migration, adhesion, contractility and clonogenic behaviour.
Gharanei S, Fishwick K, Peter Durairaj R, Jin T, Siamantouras E, Liu K-K, Straube A, Lucas ES, Weston CJ, Rantakari P, Salmi M, Jalkanen S, Brosens JJ and Tan BK
(2021)
Vascular Adhesion Protein-1 Determines the Cellular Properties of Endometrial Pericytes.
Front. Cell Dev. Biol. 8:621016.
doi: 10.3389/fcell.2020.621016
[Link]
 kinesin-3 long distance transporters
kinesin-3 long distance transporters
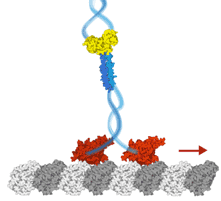
Our chapter in the kinesin superfamily handbook describes the largest subfamily, kinesin-3, a group of motors recently shown to be superprocessive and
working as cellular long-distance transporters.
Nida Siddiqui and Anne Straube (2020)
The Kinesin-3 Family: Long-Distance Transporters
in: The Kinesin Superfamily Handbook: Transporter, Creator, Destroyer, ed. Claire T. Friel, CRC Press, 2020
[Link to Book] [Open Access Chapter]
 rapid test for SARS-CoV-2
rapid test for SARS-CoV-2
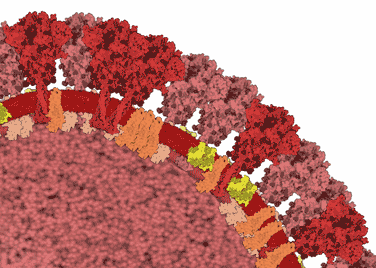
Our colleagues in the Chemistry department at Warwick developed a rapid test for SARS-CoV-2 based on the affinity of spike protein to glycans. We purified spike S1 protein from HEK cells for testing the device.
Alexander N. Baker, Sarah-Jane Richards, Collette S. Guy, Thomas R. Congdon, Muhammad Hasan, Alexander J. Zwetsloot, Angelo Gallo, Józef R. Lewandowski, Phillip J. Stansfeld, Anne Straube, Marc Walker, Simona Chessa, Giulia Pergolizzi, Simone Dedola, Robert A. Field, and Matthew I. Gibson (2020)
The SARS-COV-2 Spike Protein Binds Sialic Acids and Enables Rapid Detection in a Lateral Flow Point of Care Diagnostic Device.
ACS Central Science 6(11):2046-2052 doi: 10.1021/acscentsci.0c00855
[Link to Open Access Paper]
Alexander N. Baker, Sarah-Jane Richards, Collette S. Guy, Thomas R. Congdon, Muhammad Hasan, Alexander J. Zwetsloot, Anne Straube, Marc Walker, Simona Chessa, Giulia Pergolizzi, Simone Dedola, Robert Field, and Matthew Gibson (2020)
The SARS-COV-2 Spike Protein Binds Sialic Acids, and Enables Rapid Detection in a Lateral Flow Point of Care Diagnostic Device.
PREPRINT chemRxiv 12465680
doi: 10.26434/chemrxiv.12465680 [Link to Preprint]
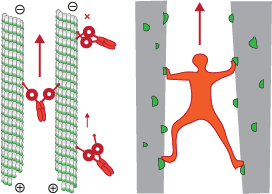 dynein activation on microtubule bundles
dynein activation on microtubule bundles
We show that human dynein is activated when it crossbridges two microtubules. Dynein polarity-sorts microtubule bundles by selectively sliding anti-parallel microtubules apart.
While single molecules of dynein are predominantly static or diffusive on single microtubules, they walks processively on the microtubule bundles they form. Their force output doubles on microtubule bundles
and we propose a model whereby the separation of the motor domains during crossbridging activates the motor.
Manas Chakraborty, Algirdas Toleikis, Nida Siddiqui, Robert A. Cross, and Anne Straube (2020)
Activation of cytoplasmic dynein through microtubule crossbridging.
PREPRINT bioRxiv 2020.04.13.038950
doi: 10.1101/2020.04.13.038950. [Link]
This preprint has been highlighted by a prelight.
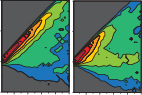
 synergy of microtubule-targeting drugs
synergy of microtubule-targeting drugs
The screening team at the Beatson Institute identified Mebendazole as a candidate to act in synergy with Docetaxel. We contributed
experiments in RPE1 and PC3 cells treated with each of the drugs individually and together to show how they affect microtubule
assembly and mitotic progression. The synergy of both drugs results frequently in mitotic catastrophe.
Linda Rushworth, Kay Hewit, Sophie Munnings-Tomes, Sukrut Somani, Daniel James, Emma Shanks, Christine Dufès, Anne Straube, Rachana Patel, and Hing Y Leung (2019)
Repurposing screen identifies mebendazole as a clinical candidate to synergise with docetaxel for prostate cancer treatment.
British Journal of Cancer.
doi: 10.1038/s41416-019-0681-5. [Link]
 phosphoregulation of microtubule binding
phosphoregulation of microtubule binding
Here we show that EML4 binds microtubules via the negatively charged E-hooks. Upon phosphorylation by mitotic kinases NEK6 and NEK7, EML4 binding to microtubules is reduced. This displacement of EML4 from microtubules reduces their stability. Dynamic microtubules are required for faithful chromosome segregation. Expressing non-phosphorylatable mutants of EML4 results in their association to microtubules throughout mitosis and spindle checkpoint activation and mitotic delay.
Adib R, Montgomery JM, Atherton J, O'Regan L, Richards MW, Straatman KR, Roth
D, Straube A, Bayliss R, Moores CA, Fry AM. (2019)
Mitotic phosphorylation by NEK6 and NEK7 reduces the microtubule affinity of EML4 to promote chromosome congression.
Sci Signal. 12(594):eaaw2939.
doi: 10.1126/scisignal.aaw2939.
[Link]
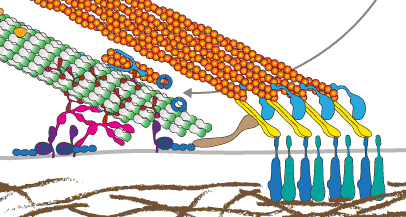 microtubules in cell migration
microtubules in cell migration
This is our contribution to the special issue on BIOCHEMICAL AND MECHANISTIC ASPECTS OF CELL MOTILITY AND MIGRATION. Our Essay summarises the functions of microtubules in cell migration including its role as directional transport tracks, signalling platforms and load-bearing elements.
Clare Garcin and Anne Straube (2019)
Microtubules in cell migration
Essays in Biochemistry 63(5): 509-520.
doi: 10.1042/EBC20190016
[Link]
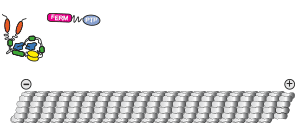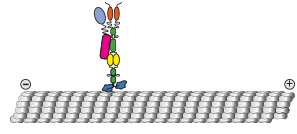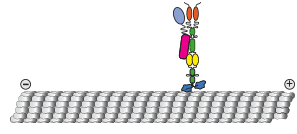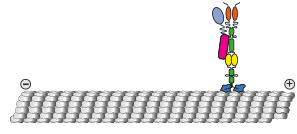 activating intracellular transport
activating intracellular transport
We show that contrary to other kinesin-3 motors, KIF1C is autoinhibited by an intramolecular interaction of its stalk with the microtubule binding surface of the motor domain. The phosphatase PTPN21 and the cargo adpater Hook3 activate KIF1C by binding to the stalk. Both activators increase the landing rate of the kinesin and are transported by the motor.
N. Siddiqui*, A.J. Zwetsloot*, A. Bachmann, D. Roth, H. Hussain, J. Brandt, I. Kaverina & A. Straube (2019)
PTPN21 and Hook3 relieve KIF1C autoinhibition and activate intracellular transport
Nature Communications, Volume 10, Article number: 2693
doi: 10.1038/s41467-019-10644-9
[Link to Paper]
Siddiqui,N., Bachmann, A., Zwetsloot,A., Hussain,H., Roth, D., Kaverina, I. and Straube, A. (2018)
Activation of intracellular transport by relieving KIF1C autoinhibition.
PREPRINT bioRxiv doi: 10.1101/488049
[Link to Preprint]
This preprint has been highlighted by a prelight.
[Link to Press Release]
 spatial EB positioning
spatial EB positioning
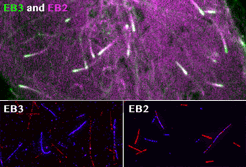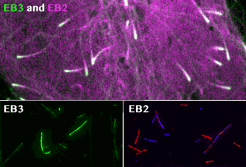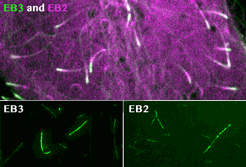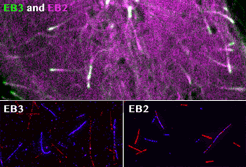
Our contribution to the special issue on reconstituting cell biology. We reconstituted the spatially distinct tip tracking of EB proteins in vitro and show that EBs sense the nucleotide state of two the beta-tubulins adjacent to their binding site. EB2 has a different nucleotide preference and prefers binding to mixed nucleotide lattices, while EB1 and EB3 need an environment of pure GTPgS to bind efficiently.
Roth, D., Fitton, B.P., Chmel, N., Wasiluk, N. and Straube, A. (2018)
Spatial positioning of EB family proteins at microtubule tips involves distinct nucleotide-dependent binding properties
Journal of Cell Science 132(4): jcs219550
doi: 10.1242/jcs.219550
[link to full pdf]
 measuring microtubule dynamics
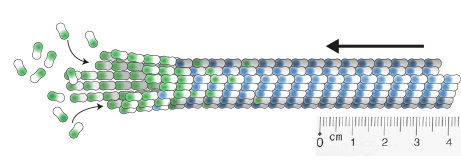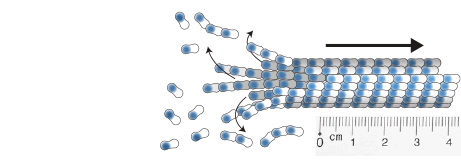
measuring microtubule dynamics
Our essay on how to measure microtubule dynamics.
Zwetsloot, A.J., Tut, G. and Straube, A. (2018)
Measuring microtubule dynamics
Essays in Biochemistry 62(6):725-735.
doi: 10.1042/EBC20180035
[link to open access article]
 kinesin-3 review
kinesin-3 review
Nida and Anne review kinesin-3 transporters, their structural characteristics, intracellular cargoes and their regulation, with particular emphasis on the autoinhibited state and mechanisms of relieving autoinhibition by cargo and adapter proteins.
Siddiqui, N. and Straube, A. (2017)
Intracellular cargo transport by kinesin-3 motors
Biochemistry (Moscow), Vol. 82, No. 7, pp. 803–815
Russian version: Biokhimiya, 2017, Vol. 82, No. 7, pp. 1047–1062
[link to open access article]
 microtubules regulating cell migration
microtubules regulating cell migration
A comprehensive book chapter in "The Microtubule Cytoskeleton" exploring the diverse functions of microtubules in cell migration and neuronal pathfinding. Co-authored with Ulrike Theisen who is studying neuronal migration in zebrafish in Braunschweig.
Ulrike Theisen and Anne Straube (2016)
Microtubules Regulate Cell Migration and Neuronal Pathfinding
in: The Microtubule Cytoskeleton, Jens Lüders (ed), pages 151-189 DOI: 10.1007/978-3-7091-1903-7_6
[Link]
 encyclopedic microtubules
encyclopedic microtubules
Anne's contribution to a major textbook with 285 chapters. My chapter contains all you ever wanted to know about microtubules and the proteins that bind and regulate them in a nutshell.
Straube, A. (2015)
Microtubules and Microtubule-Associated Proteins (MAPs)
Encyclopedia of Cell Biology, Volume 2: Organizational Cell Biology, 2016, Pages 539-547
[link] [request pdf]
 wobbly phages as nanosensors
wobbly phages as nanosensors
This paper demonstrates the use of a fluorescently-labelled and surface-anchored bacteriophage M13 to measure wall shear stress. Bacteriophage anchored on collagen-coated slides and the surface of endothelial cells were subjected to various levels of shear stress by modulating buffer flow through the imaging chamber.
Lobo, D.P. ,Wemyss, A.M., Smith, D.J., Straube, A., Betteridge, K.B., Salmon, A.H.J., Foster, R.R., Elhegni, H.E., Satchell, S.C., Little, H.A., Pacheco-Gomez, R., Simmons, M.J., Hicks, M.R., Bates, D.O., Rodger, A., Dafforn, T.R., & Arkill, K.P. (2015)
Direct detection and measurement of wall shear stress using a filamentous bio-nanoparticle.
Nano Research, doi: 10.1007/s12274-015-0831-x
[link]
 chaperoning the spindle
chaperoning the spindle
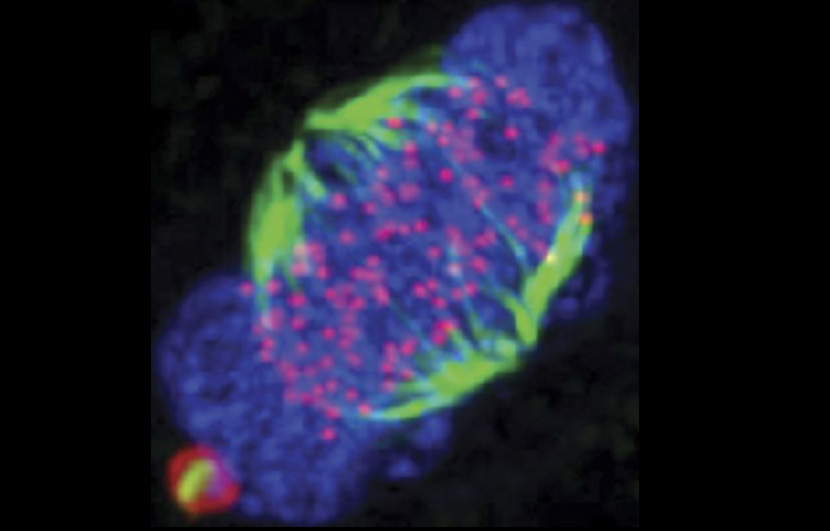
We describe Nek6-dependent spindle recruitment of Hsp70, which is required for K-fibre stability and function. Data generated in the Straube lab suggest that the weak direct interaction of Hsp70 with microtubules is not affected by the Nek6-mediated phosphorylation at T66. Thus Hsp70 possibly acts by recruiting the chTOG/TACC3/Clathrin complex to spindle microtubules.
O'Regan, L., Sampson, J., Richards, M.W., Knebel, A., Roth, D., Hood, F.E., Straube, A., Royle, S.J., Bayliss, R. & Fry, A.M. (2015)
Hsp72 is targeted to the mitotic spindle by Nek6 to promote K-fiber assembly and mitotic progression.
The Journal of Cell Biology, 209: 349-358. doi:10.1083/jcb.201409151
[link]
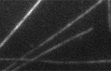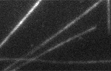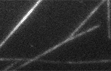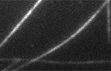 zippering microtubules
zippering microtubules
Here, we identify a previously uncharacterised isoform of microtubule-associated protein MAP4, oMAP4, as a microtubule organising factor that is crucial for myogenesis. Depletion of oMAP4 impairs cell elongation and cell-cell fusion. oMAP4 is required for paraxial microtubule organisation in muscle cells and prevents dynein- and kinesin-driven microtubule-microtubule sliding. Purified oMAP4 aligns dynamic microtubules into antiparallel bundles that withstand motor forces in vitro.
Mogessie, B., Roth, D., Rahil, Z. and Straube, A. (2015)
A novel isoform of MAP4 organises the paraxial microtubule array required for muscle cell differentiation
eLife, 4: e05697. doi:10.7554/eLife.05697.
[link] | [PDF]
 microtubule binding motif revealed
microtubule binding motif revealed
In this paper we identify the MT-binding domain in EML proteins. Daniel and Anne performed TIRF-based MT binding assays using full-length, mutant and truncation constructs of human EML1 fused to YFP. We show that EML contains a trimerisation domain that is required, but not sufficient for MT binding.
Richards, M.W., O'Regan, L., Roth, D., Montgomery, J.M., Straube, A., Fry, A.M. and Bayliss, R. (2015)
Microtubule association of EML proteins and the EML4-ALK variant 3 oncoprotein require an N-terminal trimerization domain
Biochemical Journal, 467: 529-536. doi:10.1042/BJ20150039
[link]
 transport on the go
transport on the go
Alice and Anne review the literature implicating kinesins in cell migration. It turns out we know surprisingly little about which kinesin transports which cargo to support cell polarity and directed migration. What we do know, you'll find in here.
Bachmann, A. and Straube, A. (2015)
Kinesins in cell migration
Biochem Soc Trans, 43(1): 79-83.
[PDF] | [pubmed]
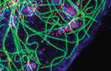 kinesin under MAP control
kinesin under MAP control
Here we show that the kinesin KIF1C is required for the de-novo formation of podosomes in vascular smooth muscle cells. We describe a pathway whereby activation of PKC results in the enrichment of CLASPs at microtubule ends, which stimulates KIF1C translocation to the cell periphery, where KIF1C localises to incipient podosome sites.
Efimova, N., Grimaldi, A., Bachmann, A., Frye, K., Zhu, X., Feoktistov, A., Straube, A. and Kaverina, I. (2014)
Podosome-regulating kinesin KIF1C translocates to the cell periphery in a CLASP-dependent manner.
Journal of Cell Science 127: 5179-5188. doi:10.1242/jcs.149633
[link]
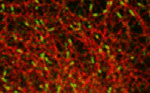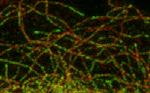 tip tracking complications
tip tracking complications
In this paper we demonstrate that CLASPs modify the EB binding site at the microtubule lattice. Ben and Daniel demonstrate the removal of the EB binding site from the microtubule lattice through CLASPs using TIRF-based in vitro reconstitution experiments.
Grimaldi, A.D., Maki, T., Fitton, B.P., Roth, D., Yampolsky, D., Davidson, M.W., Svitkina, T., Straube, A., Hayashi, I. and Kaverina, I. (2014)
CLASPs are required for proper microtubule localization of end-binding proteins.
Developmental Cell, 30, 343-352.
[link]
 exploring shapes
exploring shapes
In this paper we describe a computational framework to analyse and classify cell morphology. The approach uses a machine learning process based on the calculation of a shape similarity matrix and low-dimensional representation of cell shapes using diffusion map. We show that this approach is suitable for the highly variable cell shapes observed in migrating epithelial cells.
Jefferyes, S.D.R., Epstein, D.B.A., Straube, A., Rajpoot, N.M. (2013)
A novel framework for exploratory analysis of highly variable morphology of migrating epithelial cells.
Conf Proc IEEE Eng Med Biol Soc. 2013 Jul;2013:3463-3466, doi:10.1109/EMBC.2013.6610287
[link] [pubmed abstract]
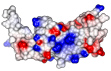 composite microtubule binders
composite microtubule binders
In this paper we demonstrate that TACC3 and clathrin form a composite microtubule binding site. Daniel and Anne did in vitro microtubule binding experiments of TACC3 and clathrin in the presence of AuroraA kinase to demonstrate the phosphorylation-dependent binding of the complex to microtubules.
Hood, F.E., Williams, S.J., Burgess, S.G., Richards, M.W., Roth, D., Straube, A., Pfuhl, M., Bayliss, R. & Royle, S.J. (2013)
Coordination of adjacent domains mediates TACC3–ch-TOG–clathrin assembly and mitotic spindle binding.
J. Cell Biol. 202:463-78
[link]
 microtubule rigidity
microtubule rigidity
In this paper we investigate the flexural rigidity of microtubules that have been stabilised by Taxol, non-hydrolysable GTP analogs or two different microtubule-associated proteins, tau and MAP4. We find that Taxol and GTPγS make microtubules more flexible, while tau and GMP-CPP make them stiffer. We show that combinations of these stabilisers do not have additive effects, but rather one stabiliser dominates the mechanical properties. MAP4 is not changing rigidity per se, but limits the variability of rigidity between microtubules.
Hawkins T.L., Sept D., Mogessie B., Straube A. and Ross J.L. (2013)
Mechanical Properties of Doubly Stabilized Microtubule Filaments.
Biophysical Journal 104: 1517-1528
[link to pdf] [pubmed abstract][request pdf]
>> see all publications here
 Clare Garcin | Postdoctoral Fellow | 2018-2022
Clare Garcin | Postdoctoral Fellow | 2018-2022
I studied microtubule organisation in differentiating muscle cells.
I was a Project Officer for Athena Swan and ED&I at Warwick Manufacturing Group and I am currently an Assistant Professor (Teaching Focussed) at Warwick Medical School in the MBChB team.
 Paula Esquivias | Postdoctoral Fellow | 2019-2021
Paula Esquivias | Postdoctoral Fellow | 2019-2021
I studied microtubule-associated protein 4.
I am currently a bioscience lead at The Rosalind Franklin Laboratory in Leamington.
 Helena Coker | Imaging Specialist | 2017-2021
Helena Coker | Imaging Specialist | 2017-2021
I setup and run the Lattice Light Sheet microscope and run the LLSM visitor programme.
I am currently an imaging specialist in Oxford.
 Jack Chen | Postdoctoral Fellow | 2017-2020
Jack Chen | Postdoctoral Fellow | 2017-2020
I studied microtubule tip tracking transport using biochemical in vitro reconstitution assays.
I am currently a scientist at Oxford Biomedica.
 Nida Siddiqui | Postdoctoral Fellow | 2018-2019
Nida Siddiqui | Postdoctoral Fellow | 2018-2019
I studied Kif1C activation and force generation. I published two preprints, that were subsequently published in Nature Communications and Current Biology, one review and one book chapter, all as first author.
I am currently a Senior Scientist at AstraZeneca in Cambridge. [LinkedIn]
 Manas Chakraborty | Postdoctoral Fellow | 2016-2019
Manas Chakraborty | Postdoctoral Fellow | 2016-2019
I studied microtubule self-organisation using biochemical in vitro reconstitution assays.
I am currently back in India and looking for faculty positions.
 Mohammed Abdelsamea | Postdoctoral Fellow | 2018
Mohammed Abdelsamea | Postdoctoral Fellow | 2018
I developed image analysis software.
I moved on to bea postdoc at Nottingham University, a senor lecturer at Birmingham City University and I am now a senior lecturer at University of Exeter. [web]
 Gaëlle Letort | Software Developer | 2016
Gaëlle Letort | Software Developer | 2016
I developed image analysis routines to segment and measure microtubules in 3D superresolution images of human skeletal muscle cells.
I moved on to be a postdoc at the Institute Curie in Paris, France and I am now a research engineer at Institute Pasteur. [LinkedIn]
I have a PhD in Computer Science and experience as a software developer with a specialty in human-computer interaction. I developed user-friendly software packages to analyse microtubule dynamics and cell shape.
I moved on to be a senior scientific developer at zfx-innovation in Gargazzone, Italy. and I am now Head of Quality Assurance at Zeiss Digital Innovation [LinkedIn]
 Ulrike Theisen | postdoctoral research fellow | 2008 - 2012
Ulrike Theisen | postdoctoral research fellow | 2008 - 2012
Kinesin motors perform the long-distance transport towards the plus end of microtubules. Thus kinesins are likely to be responsible for delivering polarity factors to the cell edges. I have identified Kif1C as the transporter for rearward transport of integrins in migrating cells and shown that this transport supports tail stability and rear drag, a mechanism important for directionally persistent cell migration in the absence of external guidance cues (Theisen et al., Dev Cell 2012).
I am currently a Marie Curie Fellow at Technische Universität Braunschweig and study neuronal migration in Zebrafish. [web]
 Mike Downey | research associate | 2012-2013
Mike Downey | research associate | 2012-2013
I am a chemist and computer scientist by training and applied my expertise in semiautomatic image analysis to study how dynamic microtubules regulate cell shape and directional-persistent migration.
I moved on to be a postdoc in Till Bretschneider's group at the Warwick Systems Biology Centre and develop software tools for studying 3D cell shape dynamics.
I am now a software developer / data analyst in Birmingham.
 Lewis Mosby |PhD student | 2017-2021
Lewis Mosby |PhD student | 2017-2021
I derived beautiful equations to model tip tracking transport.
I was an IAS Early Career Fellow at Warwick and I am currently a postdoc with Zena Hadjivasiliou at the Francis Crick Institute.
 Alexander Zwetsloot | PhD student | 2017-2021
Alexander Zwetsloot | PhD student | 2017-2021
I reconstituted bidirectional transport complexes with KIF1C and dynein (Abid Ali, Zwetsloot et al. Nature Structural & Molecular Biology 2025). I also discovered that Hook3 activates and co-transports with KIF1C (Siddiqui, Zwetsloot et al. Nature Communications 2019), I contributed to
the development of a rapid detection device for Cov-SARS-2 (Baker et al. ACS Central Science 2020) and to the analysis of HSP patient mutations in KIF1C (Siddiqui et al. bioRxiv 2021), and wrote a review (Zwetsloot, Tut & Straube, Essays in Biochemistry 2018).
I am currently a scientific investigator at GSK. [LinkedIn]
 Jonathan Brandt | PhD student | 2016-2020
Jonathan Brandt | PhD student | 2016-2020
I studied the changes in the host cytoskeleton upon infection with Marek's disease virus. In particular, I focussed on the non-catalytic role of Us3 kinase on actin dynamics.
 Gokhan Tut | PhD student | 2015 - 2018
Gokhan Tut | PhD student | 2015 - 2018
I studied the intracellular transport of Marek's Disease Virus.
I am currently a postdoc with Paul Moss at the Institute of Immunology and Immunotherapy, University of Birmingham.
 Nida Siddiqui | PhD student | 2014 - 2018
Nida Siddiqui | PhD student | 2014 - 2018
I studied the autoinhibition mechanisms of Kif1C and its activation by PTPN21 (Siddiqui et al. Nat Comms 2019).
I stayed on as a postdoc in the Straube lab. See above what happened next.
 Alice Bachmann | PhD student | 2013 - 2017
Alice Bachmann | PhD student | 2013 - 2017
I studied the mechanisms by which the kinesin Kif1C promotes the formation of podosomes in vascular smooth muscle cells (Efimova et al. J Cell Sci 2014) and identified PTPN21 as an activator of KIF1C-dependent intracellular transport (Siddiqui et al. Nat Comms 2019).
I am currently working for BioRad in Oxford.
 Ben Fitton | PhD student | 2011 - 2016
Ben Fitton | PhD student | 2011 - 2016
Microtubule dynamics is characterised by phases of continuous growth and shrinkage and stochastic switches between these phases. I studied the transitions between growth and shrinkage phases, catastrophes and rescues, in detail using in vitro reconstitution of microtubule assembly/disassembly in the presence of EB proteins to detect the GTP cap.
I am currently on a civil service fast track apprenticeship applying my mathematical modelling skills to new kinds of problems.
 Sam Jefferyes | PhD student | 2011-2015
Sam Jefferyes | PhD student | 2011-2015
Migrating cells adopt a variety of different shapes, which are determined by the cytoskeleton in response to the extracellular substrate. I developed machine learning algorithms for the unbiased quantification of cell morphology. I further analysed cell shape changes of migrating cells and developed models to predict cell behaviour from cell shape data alone.
I am currently software developer at quantumkdb in London - Glasgow - Ireland. [web]
 Binyam Mogessie | PhD student
| 2007-2011
Binyam Mogessie | PhD student
| 2007-2011
I investigated the role of MAP4 in cell division and muscle cell differentiation. I revealed a role for MAP4 in limiting force generation by cortical dynein in mitotic cells. This is important for spindle positioning (Samora, Mogessie et al., Nature Cell Biol 2011). I have also identified a novel MAP4 isoform required for the formation of antiparallel microtubule arrangement in differentiating muscle cells (Mogessie et al. eLife 2015).
I was a postdoc with Melina Schuh at the MRC Laboratory of Molecular Biology in Cambridge and a group leader at the University of Bristol. I am now an assistant professor at Yale [web]
 Olivier Tardy | Research Assistant | 2022
Olivier Tardy | Research Assistant | 2022
I purified dynein.
 Isaac Gardiner | part-time technician | 2022
Isaac Gardiner | part-time technician | 2022
I learned how to express proteins in insect cells.
 Hannah Gay | part-time technician | 2016
Hannah Gay | part-time technician | 2016
I tried to generate homologous repair constructs and guideRNAs for CRISPR/Cas9-mediated labelling of endogenous cytoskeleton proteins with fluorescent markers.
I am now a PhD student in Birmingham.
 Ana Clark (now Wallis) | part-time technician | 2015
Ana Clark (now Wallis) | part-time technician | 2015
I performed 2D/3D cell migration assays with human breast cancer cells.
I have since completed a PhD in Oxford and I am now working in science communication at St John's College and the Natural History Museum in Oxford.
 Zoe Redshaw | part-time technician | 2014
Zoe Redshaw | part-time technician | 2014
I selected stable human breast cancer cell lines.
I am now a postdoc in Nottingham.
 Cyntia Fernandez Cuesta| Rotation student Midlands Integrative Biosciences Training Programme | 2020
Cyntia Fernandez Cuesta| Rotation student Midlands Integrative Biosciences Training Programme | 2020
I studied Us3 kinase of Marek's disease virus.
I am now undertaking my second miniproject and then my PhD with Aga Gambus at Birmingham University.
 Christian Clarke| Rotation student Midlands Integrative Biosciences Training Programme | 2020
Christian Clarke| Rotation student Midlands Integrative Biosciences Training Programme | 2020
I studied the transport of Marek's disease virus.
I am now undertaking my second miniproject and then my PhD at Leicester.
 Hannah Smith| Rotation student Midlands Integrative Biosciences Training Programme | 2019
Hannah Smith| Rotation student Midlands Integrative Biosciences Training Programme | 2019
I studied the transport of Marek's disease virus.
I am now undertaking my PhD at Leicester.
 Alice Haworth| Rotation student Midlands Integrative Biosciences Training Programme | 2017
Alice Haworth| Rotation student Midlands Integrative Biosciences Training Programme | 2017
I studied KIF15 localisation throughout the cell cycle.
I am now undertaking my PhD at Leicester.
 Ainur Kakpenova | MSc by research student | 2014 - 2015
Ainur Kakpenova | MSc by research student | 2014 - 2015
I studied spatial microtubule dynamics regulation in migrating cells.
I am now back at Nazarbayev University in Kazakhstan.
 Jacopo Credi | Erasmus Mundus MSc in Complex Systems Science | 2015
Jacopo Credi | Erasmus Mundus MSc in Complex Systems Science | 2015
I studied swarming behaviour of migrating cancer cells.
I am now undertaking the second year of my MSc at Ecole Polytechnique in Paris.
 Alina Finch| Rotation student Midlands Integrative Biosciences Training Programme | 2015
Alina Finch| Rotation student Midlands Integrative Biosciences Training Programme | 2015
I studied kinesin functions in cell migration.
I am now undertaking my PhD at Leicester.
 Alexandra Matthews| Rotation student Midlands Integrative Biosciences Training Programme | 2015
Alexandra Matthews| Rotation student Midlands Integrative Biosciences Training Programme | 2015
I studied kinesin functions in cell migration.
I am now undertaking my PhD at Birmingham.
 Justyna Szyroka | Rotation student Midlands Integrative Biosciences Training Programme | 2015
Justyna Szyroka | Rotation student Midlands Integrative Biosciences Training Programme | 2015
I studied changes in cell shape and motility of chicken embryonic fibroblasts upon infection with Marek's disease virus.
I am now undertaking my PhD at Warwick.
 Diana Sifuentes Munch | Erasmus Mundus MSc in Complex Systems Science | 2014
Diana Sifuentes Munch | Erasmus Mundus MSc in Complex Systems Science | 2014
I analysed correlations between cell shape and cell migration behaviour.
I am now undertaking the second year of my MSc at Ecole Polytechnique in Paris.
 Jessica Talbot | MSc in Complexity Science | 2014
Jessica Talbot | MSc in Complexity Science | 2014
I worked with Stefan Grosskinsky (Warwick Complexity Centre) to simulate cell migration and division on a 2D surface with a sequential lattice model.
I am now undertaking the second miniproject for my MSc.
 Lewis Baker | MSc student MOAC | 2014
Lewis Baker | MSc student MOAC | 2014
I studied actin dynamics in migrating cells and the correlation with growing microtubule ends in proximity of the cell cortex.
I am now a PhD student.
 Scott Clarke | MSc student MOAC| 2014
Scott Clarke | MSc student MOAC| 2014
I purified tubulin from human cells.
I am now a PhD student with Mohan Balasubramanian.
 Rozita Adib | Rotation student Midlands Integrative Biosciences Training Programme| 2014
Rozita Adib | Rotation student Midlands Integrative Biosciences Training Programme| 2014
I studied the effect of EML1 depletion on microtubule asymmetry in migrating cells.
I am now a PhD student with Andrew Fry at Leicester University.
 Harriet Bell | MSc student Scientific Research and Communication| 2013
Harriet Bell | MSc student Scientific Research and Communication| 2013
I studied the kinesin Kif1C and its interactors to find out how Kif1C transports vesicles along microtubules. In addition to my research I worked part time with the children's cancer team at the Cancer Research UK Clinical Trials Unit, University of Birmingham.
 Nikita Nicholls | MSc student Systems Biology | 2012
Nikita Nicholls | MSc student Systems Biology | 2012
I studied kinesins in spindle positioning.
I am now a PhD student in the Midlands Integrative Biosciences Training Partnership.
 Sarah Cosgriff | MSc student Systems Biology | 2011
Sarah Cosgriff | MSc student Systems Biology | 2011
I studied cell shape changes and actin dynamics at the cell front in migrating cells.
I am now administrator at STEMNET and work at the University of Birmingham's STEM Centre to coordinate work with schools in Birmingham and Solihull.
 Robert Lockley | MSc student Systems Biology | 2011
Robert Lockley | MSc student Systems Biology | 2011
I studied the directionality of microtubule assembly in migrating cells.
I am now a PhD student with Graham Ladds and Till Brettschneider at the University of Warwick.
 Charlotte Carroll | rotation student Life Sciences PhD | 2011
Charlotte Carroll | rotation student Life Sciences PhD | 2011
I studied the competition between EB1, EB2 and EB3 at the microtubule plus end in vitro.
I am now a PhD student with Lorenzo Frigerio at the University of Warwick.
 Harold Moyse | MSc student Systems Biology | 2011
Harold Moyse | MSc student Systems Biology | 2011
I studied the migration of differentiating muscle cells.
I am now a PhD student with Neils Evans and Daniel Zehnder the University of Warwick.
 Rachel Sheldon | MSc student MOAC | 2011
Rachel Sheldon | MSc student MOAC | 2011
I studied the microtubule binding properties of EB proteins.
I am now a PhD student with Hugo van den Berg, Anatoly Shmygol and Andy Blanks at the University of Warwick.
 Ali Rasooli-Nejad
| MSc student Systems Biology | 2010
Ali Rasooli-Nejad
| MSc student Systems Biology | 2010
I studied cell behaviour during contact inhibition of locomotion.
I am now a PhD student with Yuriy Pankratov at the University of Warwick.
 Aida Hassan | Lister Institute summer student | 2020
Aida Hassan | Lister Institute summer student | 2020
I analysed single molecule kinetics of EB proteins.
I am now completing my degree at the University of Nottingham.
 Julia Schander | Lister Institute summer student | 2019
Julia Schander | Lister Institute summer student | 2019
I studied kinesin-3 motors.
I am now completing my degree in process engineering at the TU Dresden.
 Samia Mohammed | BSCB summer student | 2019
Samia Mohammed | BSCB summer student | 2019
I studied microtubule bundling by dynein.
I am now completing my degree in biomedical sciences at Birmingham City University.
 Elly Straube | Lister Institute summer student | 2018
Elly Straube | Lister Institute summer student | 2018
I engineered and characterised inhibitable Kif1C constructs.
I am now completing my degree in process engineering at the TU Dresden.
 Meghana Kumar | Lister Institute summer student | 2016
Meghana Kumar | Lister Institute summer student | 2016
I studied the regulation of microtubule dynamics by Kif1C.
I was an MRes student at KCL and I am now working with Monica Gotta at the University of Geneva.
 Nida Siddiqui | Lister Institute summer student | 2014
Nida Siddiqui | Lister Institute summer student | 2014
I isolated tubulin from different sources rich and low in posttranslational modifications.
I was a PhD student and then a postdoc in the Straube lab.
 Luke Edwards | summer student | 2013
Luke Edwards | summer student | 2013
I spent my summer in the cold room to establish single-step purification of human tubulin using an affinity column with TOG domains.
I have since completed a BSc Cellular and Molecular Medicine at Bristol University and I am now a support worker at Care UK.
 Zainab Rahil | summer student | 2012
Zainab Rahil | summer student | 2012
I analysed microtubule directionality and orderliness in muscle cells (Mogessie et al. eLife 2015).
I am now a PhD student in the Leckband group at the University of Illinois.
 Natalia Wasiluk | Erasmus student | 2010
Natalia Wasiluk | Erasmus student | 2010
I spent a training internship in my final year as an undergraduate in the Straube lab and established TIRF-based in vitro microtubule plus end tracking assays using EB1, EB2 and EB3 (Roth et al., J Cell Sci 2019).
I am now working in the University of Cambridge Metabolic Research Laboratories.
 Agnieszka Skalecka | summer student | 2008
Agnieszka Skalecka | summer student | 2008
As a summer student at the Marie Curie Research Institute, I characterised EB3-specific interacting proteins.
I have since completed a PhD student in Jacek Jaworski's lab at the IIMCB in Warsaw, Poland [web] and I am now a postdoc at McGill University in Canada
Investigator Award in Science | Wellcome Trust
How cargo transporters switch into microtubule organisers
£1,453,634 - 2023 to 2027
Investigator Award in Science | Wellcome Trust
Self-organisation of acentrosomal microtubule arrays
£943,108 - 2016 to 2022
Project Grant | Leverhulme Trust
How microtubule plus tip trackers couple polymer assembly to cargo transport
Co-Applicants: Marco Polin and Robert Cross
£256,434 - 2017 to 2021
Lister Research Prize | The Lister Institute of Preventive Medicine
Microtubule-actin crosstalk in cell migration
£200,000 - 2014 to 2019
Warwick Quantitative Biomedicine Programme Seed Fund | Wellcome Trust / University of Warwick
Microtubule control of cell and tissue morphology
£25,000 - 2015 to 2016
PhD studentship | British Heart Foundation
Kinesin-control of podosome formation of vascular smooth muscle cells
£105,584 - 2013 to 2016
Programme Grant | Marie Curie Cancer Care
Mechanisms of directional cell migration and differentiation
£630,000 - 2009 to 2013
Core Funding | Marie Curie Cancer Care
Cytoskeletal Organisation Laboratory
2007 to 2009
Brazil Partnership Fund | Santander / University of Warwick
Collaboration with Deborah Schechtman - Phosphoregulation of tubulin assembly
£8,700 - 2014 to 2016
Collaboration Fund | EPSRC Grand Challenge network "Understanding the Physics of Life"
Collaboration with Massimo Antognozzi - Lateral Motion Force Microscopy of microtubule dynamics
£2,405 - 2014
Research Development Fund | University of Warwick
Breast cancer cell migration
£9,000 - 2014 to 2015
Strategic Partnership Fund | University of Warwick
Collaboration with Irina Kaverina (Vanderbilt) - Microtubules in directional migration
£10,000 - 2011 to 2012
Joint Warwick-QMUL Awards for Collaborative Research
Collaboration with Ann Wheeler (QMUL) - Superresolution imaging of +TIPs
£29,123 - 2012 to 2013
>> Educational magazine contribution
Biological Sciences Review is a magazine for A level students. Find us in Volume 36 Issue 2, where Anne explains how cells change their shape.
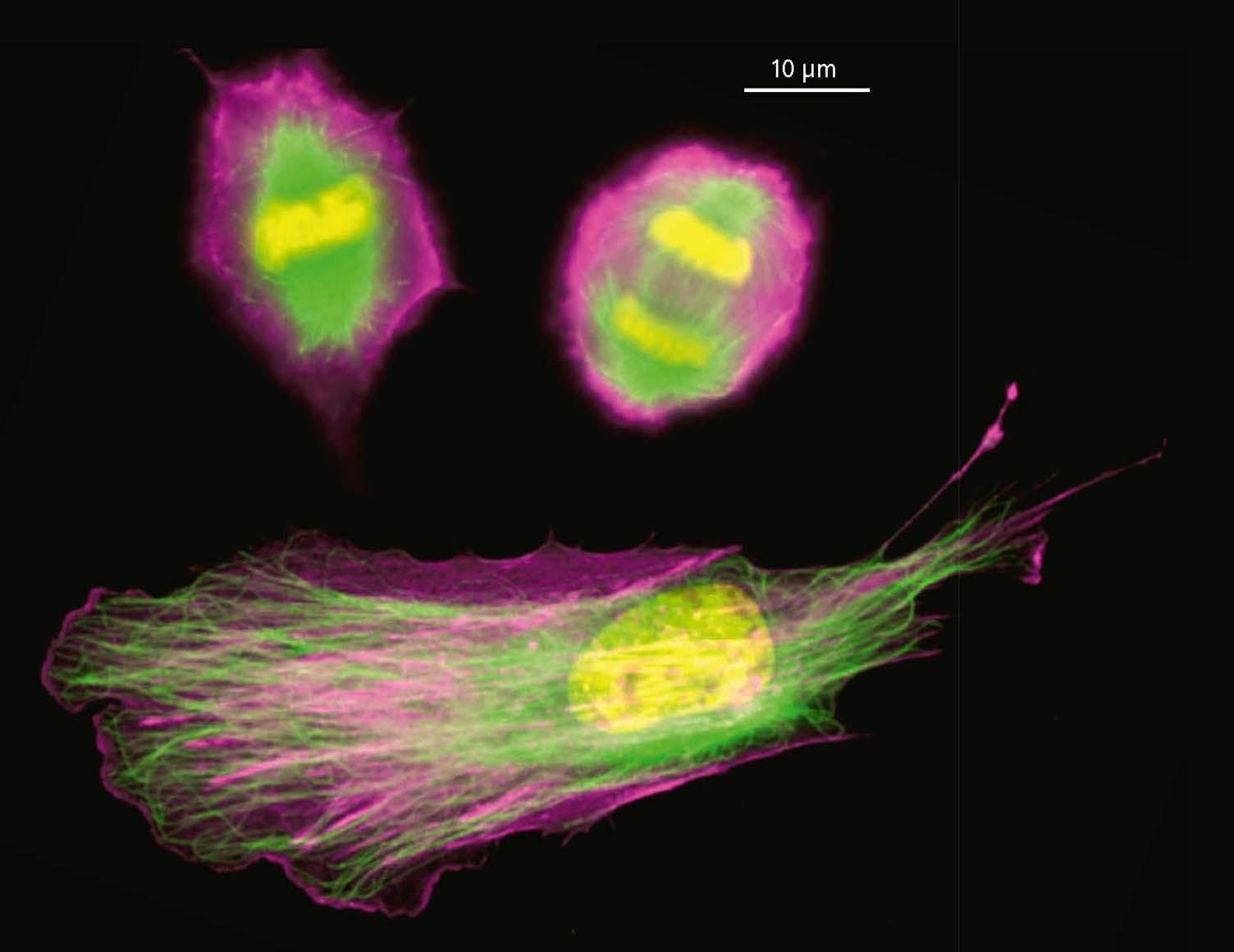 top | Human retinal epithelial cells undergoing mitosis with cytoskeletal elements and DNA stained (DNA is yellow, actin is pink, microtubules are green.
top | Human retinal epithelial cells undergoing mitosis with cytoskeletal elements and DNA stained (DNA is yellow, actin is pink, microtubules are green.
>> Microtubule brick modelling
Our school outreach programme includes building models of intracellular molecular machines from construction toy bricks and we are also happy to deliver hands-on DNA isolation workshops to primary or secondary schools. If you are interested in hosting a visit from us, please do get in touch.
 top | Second generation brick model of a polymerising microtubule end, alpha-tubulin in black, GTP-bound beta-tubulin in yellow, GDP-bound beta-tubulin in red.
top | Second generation brick model of a polymerising microtubule end, alpha-tubulin in black, GTP-bound beta-tubulin in yellow, GDP-bound beta-tubulin in red.
 left | My son has been testing my brick collection and we built: a DNA double helix, a mitotic cell picture and a viral capsid. So it's not just microtubules...
left | My son has been testing my brick collection and we built: a DNA double helix, a mitotic cell picture and a viral capsid. So it's not just microtubules...
>> Pictures from the exhibition of our Art & Science project "The Fox Got You" with Françoise Sergy in the Oxford Botanic Garden from 4th July to 31st August 2015.
 left | Anne explains the wonders of microtubule research and how microtubule-targeting drugs work - at the public event during the exhibition at the Oxford Botanic Garden.
left | Anne explains the wonders of microtubule research and how microtubule-targeting drugs work - at the public event during the exhibition at the Oxford Botanic Garden.
below | Impressions from the exhibition describing 5 medicinal plants and their use in research and the clinic.
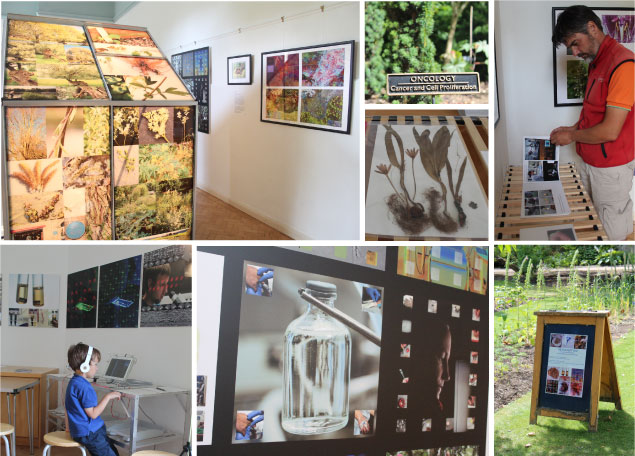

Please visit the website documenting the entire project including the laboratory visits. There is a lot to learn how to grow the plants that make important medicines and to be impressed by the MRC units giant LEGO enzyme!
more coming soon...
In the future you will find here:
>> Instructions how to build a microtubule from LEGO bricks and images of our first and second generation microtubule models.
>> Plain english explanations of our research projects.

There are a number of opportunities to join us. We are an interdisciplinary and international group, united by the love of science, our famous cakeclub and any other eggcellent morale boosting activities.
Two Wellcome Trust-funded Positions are available immediately to work on an interdisciplinary project ranging from live cell imaging to single molecule biophysics of molecular motors.
Postdoctoral research fellow role with application deadline 15th August 2024. Please find application link and further information here.
Research Assistant role with application deadline 4th September 2024. Please find application link and further information here.
One summer studentship for 10 weeks is available in my lab each summer. You will work on a quantitative imaging project and receive a stipend of £235 per week. If you cannot imagine a more exciting summer than one spent in the lab, please send me a CV, a personal statement and one academic reference by January each year.
PhD students and postdocs
We are always looking for excellent students and postdocs to apply for scholarships and fellowships. Bridging funds for 1 year and
mentoring support are available to strong candidates with matching research interests who want to apply for fellowships. If you are
motivated to work in an interdisciplinary environment, have a background in biology, physics, maths or computer
science and are enthusiastic about our research, then please get in touch with your CV and a brief statement of research interests.
Postdoc positions are available. Please apply to any advertised positions or just get in touch directly to discuss the projects.


 motor co-dependence mechanisms
motor co-dependence mechanisms surfing the end
surfing the end weak motors cause neuronal disease
weak motors cause neuronal disease end binding proteins
end binding proteins endometrial pericyte migration
endometrial pericyte migration kinesin-3 long distance transporters
kinesin-3 long distance transporters rapid test for SARS-CoV-2
rapid test for SARS-CoV-2 dynein activation on microtubule bundles
dynein activation on microtubule bundles synergy of microtubule-targeting drugs
synergy of microtubule-targeting drugs phosphoregulation of microtubule binding
phosphoregulation of microtubule binding microtubules in cell migration
microtubules in cell migration activating intracellular transport
activating intracellular transport spatial EB positioning
spatial EB positioning measuring microtubule dynamics
measuring microtubule dynamics kinesin-3 review
kinesin-3 review microtubules regulating cell migration
microtubules regulating cell migration encyclopedic microtubules
encyclopedic microtubules wobbly phages as nanosensors
wobbly phages as nanosensors chaperoning the spindle
chaperoning the spindle zippering microtubules
zippering microtubules microtubule binding motif revealed
microtubule binding motif revealed transport on the go
transport on the go kinesin under MAP control
kinesin under MAP control tip tracking complications
tip tracking complications exploring shapes
exploring shapes composite microtubule binders
composite microtubule binders microtubule rigidity
microtubule rigidity

 Research summary
Research summary Anne Straube
Anne Straube  Daniel Roth
Daniel Roth  David Corcoran
David Corcoran  Lokesh Kumar
Lokesh Kumar  Jessica Bithell
Jessica Bithell  Sareeta Bagri
Sareeta Bagri  Mahir Taher
Mahir Taher  Katie Brooks
Katie Brooks 
 Clare Garcin
Clare Garcin  Paula Esquivias
Paula Esquivias  Helena Coker
Helena Coker  Jack Chen
Jack Chen  Nida Siddiqui
Nida Siddiqui  Manas Chakraborty
Manas Chakraborty  Mohammed Abdelsamea
Mohammed Abdelsamea  Gaëlle Letort
Gaëlle Letort  Rose Gostner
Rose Gostner  Ulrike Theisen
Ulrike Theisen  Mike Downey
Mike Downey 
 Lewis Mosby
Lewis Mosby  Alexander Zwetsloot
Alexander Zwetsloot  Jonathan Brandt
Jonathan Brandt  Gokhan Tut
Gokhan Tut  Alice Bachmann
Alice Bachmann  Ben Fitton
Ben Fitton  Sam Jefferyes
Sam Jefferyes  Binyam Mogessie
Binyam Mogessie 
 Olivier Tardy
Olivier Tardy  Isaac Gardiner
Isaac Gardiner  Hannah Gay
Hannah Gay  Ana Clark (now Wallis)
Ana Clark (now Wallis)  Zoe Redshaw
Zoe Redshaw 
 Cyntia Fernandez Cuesta
Cyntia Fernandez Cuesta Christian Clarke
Christian Clarke Hannah Smith
Hannah Smith Alice Haworth
Alice Haworth Ainur Kakpenova
Ainur Kakpenova  Jacopo Credi
Jacopo Credi  Alina Finch
Alina Finch Alexandra Matthews
Alexandra Matthews Justyna Szyroka
Justyna Szyroka  Diana Sifuentes Munch
Diana Sifuentes Munch  Jessica Talbot
Jessica Talbot  Lewis Baker
Lewis Baker  Scott Clarke
Scott Clarke  Rozita Adib
Rozita Adib  Harriet Bell
Harriet Bell  Nikita Nicholls
Nikita Nicholls  Sarah Cosgriff
Sarah Cosgriff  Robert Lockley
Robert Lockley  Charlotte Carroll
Charlotte Carroll  Harold Moyse
Harold Moyse  Rachel Sheldon
Rachel Sheldon  Ali Rasooli-Nejad
Ali Rasooli-Nejad

 Aida Hassan
Aida Hassan  Julia Schander
Julia Schander  Samia Mohammed
Samia Mohammed  Elly Straube
Elly Straube  Meghana Kumar
Meghana Kumar  Luke Edwards
Luke Edwards  Zainab Rahil
Zainab Rahil  Natalia Wasiluk
Natalia Wasiluk  Agnieszka Skalecka
Agnieszka Skalecka 
 Andrew Carter
Andrew Carter  Andrew McAinsh
Andrew McAinsh  Jenny Ross
Jenny Ross 
 Andrew Fry and Richard Bayliss
Andrew Fry and Richard Bayliss  Irina Kaverina
Irina Kaverina  Ekkehard Straube
Ekkehard Straube  Nasir Rajpoot
Nasir Rajpoot  Steve Royle
Steve Royle